RCANE: A Deep Learning Algorithm for Whole-genome Pan-Cancer Somatic Copy Number Aberration Prediction using RNA-seq Data
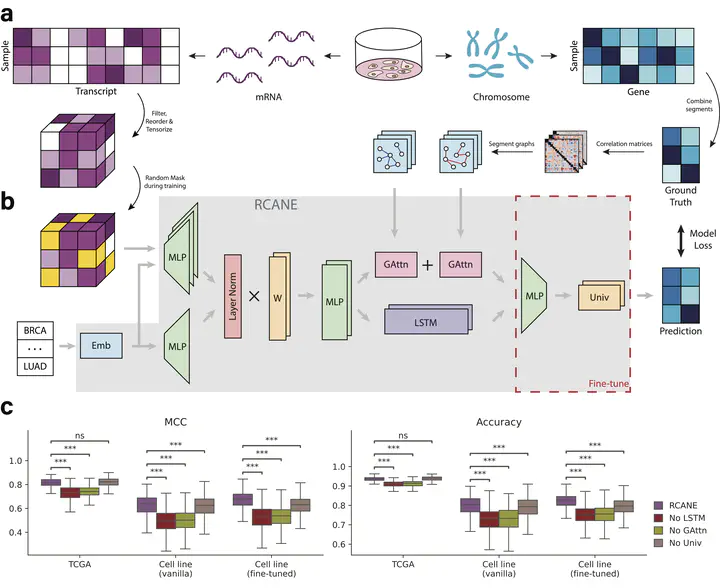 RCANE
RCANEAbstract
Transcriptome sequencing (RNA-seq) of cancers is widely employed in cancer research to investigate gene expression patterns and their role in disease progression. Somatic copy-number aberrations (SCNAs)—critical genomic drivers of tumorigenesis—can also be inferred directly from RNA-seq, yielding a “two-for-one” return of quantitative expression measures plus structural-variation calls at a fraction of the cost of separate DNA assays. Here, we present RCANE, a deep-learning framework that predicts genome-wide SCNAs across diverse cancer types using only RNA-seq data. Trained on The Cancer Genome Atlas (TCGA) and DepMap cell-line cohorts, RCANE consistently outperforms existing approaches, delivering a scalable, robust solution for improving somatic copy-number aberration profiling in cancer diagnostics and therapeutic decision-making.